Several methods are currently available for the analysis of antigenic epitopes: X-ray diffraction, phage presentation of random peptide libraries, LC-MS analysis, and 3D-EM analysis. The application of 3D-EM in biology has not only remained in a simple and intuitive description, but also has carried out a three-dimensional study from qualitative to quantitative, from plane to space. This is of great significance for understanding the spatial structure of biological macromolecules and their relationship with function. Among them, the method of X-ray crystal diffraction is the most accurate method, and the method of antigen-antibody co-crystallization can be accurate to the atomic level.
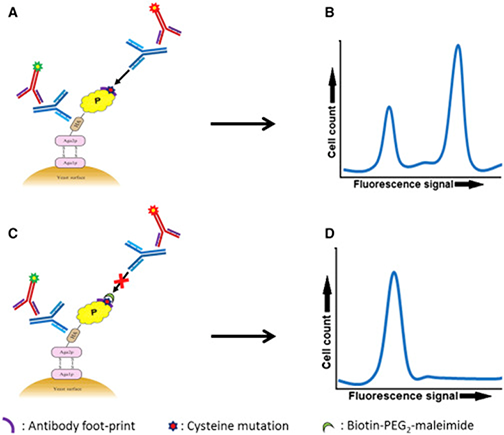 Fig.1 Schematic Outline of the Epitope Mapping Approach (Najar, et al. 2017)
Fig.1 Schematic Outline of the Epitope Mapping Approach (Najar, et al. 2017)
HDX-MS is a method for studying protein-ligand interactions and protein-protein interactions. The basic principle of this method is to immerse proteins in heavy water. The hydrogen on the surface of the protein in close contact with heavy water is faster than the hydrogen exchange rate inside the protein. Once the two proteins bind to each other, the solvent cannot reach the interface of its interaction. This principle is reflected in the hydroquinone exchange kinetics, and the amount of ruthenium contained in the antigen when bound or not bound to the monoclonal antibody can be obtained by mass spectrometry.
X-ray-resolved monoclonal antibody-antigen complexes are the most reliable method for analyzing epitope recognition by monoclonal antibodies. The method is based on the intensity and angle measurement of the X-ray beam of the monoclonal antibody-antigen eutectic diffraction, and the bioinformatics method is used to determine the atomic coordinates. As a result, the three-dimensional structure of the complex is completely visualized.
N62 is a monoclonal antibody to Francisella tularensis. The X-ray crystal structure of the Fab segment of monoclonal antibody N62 shows that the antigen-binding region is composed of aromatic amino acids, and the aromatic amino acid forms a small recess containing no more than 4/3 sugar residues, indicating that N62 is mainly The Qui4NFm residue at the non-reducing end of the Franciamycin T antigen binds.
Mnt C is a Staphylococcus aureus transporter. GRIBENKO et al. used X-ray crystallography to analyze the complex of Mnt C and its monoclonal antibody 305-78-7. It is shown that 305-78-7 binds to the C-terminal leaflet of Mnt C, and α-helixα8 and β-sheetβ8 constitute the major interface of the antigen-antibody. In addition, E247 of Mnt C may form a salt bridge with K58 on the heavy chain of the antibody.
Creative Peptides provides a comprehensive series of Epitope Mapping Services to fulfill customers' demand. Our Creative MapTM Platform is developed to map various epitopes flexibly. Based on the three-dimensional electron microscopy technology, the epitope analysis technique requires only a small amount of antigen/antigen complex to be completed within quickly.
High Resolution Conformational Epitope Mapping is a method used to identify and analyze the three-dimensional structure of antigenic epitopes recognized by antibodies. It provides insights into both linear and conformational epitopes, enhancing our understanding of protein-antibody interactions at an atomic level.
Hydrogen Deuterium Exchange Mass Spectrometry (HDX-MS) is used to study protein-ligand and protein-protein interactions. The method involves soaking proteins in heavy water to track hydrogen exchange rates, revealing the interaction sites between antibodies and antigens at the molecular level.
X-ray diffraction, especially when applied to monoclonal antibody-antigen co-crystallization, provides the most accurate three-dimensional structure of antigen-antibody complexes. This method allows for atomic-level visualization of the binding interface, offering precise details about epitope recognition.
3D-Electron Microscopy (3D-EM) offers a high-resolution approach to study the spatial structure of biological macromolecules. This method allows for the visualization of epitope structures and their interactions with antibodies in a three-dimensional context, providing a detailed view of protein functionality.
X-ray-resolved complexes offer the most reliable method for analyzing epitope recognition. By measuring the intensity and angle of X-ray beams, the method generates accurate atomic coordinates, enabling a complete visualization of the antibody-antigen interaction at the atomic level.
Our Creative MapTM Platform uses advanced 3D-EM technology to map epitopes efficiently, even with small amounts of antigen or antigen complexes. This platform provides fast and precise results, tailored to meet various research needs in epitope analysis.
The X-ray crystal structures of monoclonal antibodies like N62 and Mnt C provide detailed insights into the antibody-antigen interface. These structures help identify crucial binding residues, such as aromatic amino acids and specific salt bridges, which are essential for understanding how antibodies interact with antigens.

References