Peptide Mass Fingerprinting (PMF) is a powerful analytical technique used in proteomics to identify proteins based on their peptide mass profiles. It involves digesting proteins into peptides using enzymes like trypsin, separating these peptides using techniques such as mass spectrometry, and then matching the observed peptide masses with theoretical masses derived from protein sequence databases. Creative Peptides offers comprehensive services in peptide synthesis and analysis, including PMF, utilizing state-of-the-art equipment and expertise to support precise protein identification and characterization needs. Their commitment to quality and innovation makes them a reliable partner for researchers and industries seeking accurate proteomic solutions.
Peptide Mass Fingerprinting (PMF) assay is a method for mass spectrometric analysis of a mixture of polypeptides obtained after proteolysis or degradation. The peptide obtained by mass spectrometry is compared with the theoretical peptide of the protein in the polypeptide protein database to determine whether the measured protein is known or unknown. Since different proteins have different amino acid sequences, the peptides obtained from different proteins have fingerprint characteristics. The absolute mass of each protein peptide is theoretically calculated to find the best match compared to the mass of the unknown protein peptide. A variety of proteomes have been studied using peptide fingerprinting methods.
The advantage of PMF is that if the protein is present in the protein database, only the mass of the protein peptide is known, and it takes no time to perform DEVONO sequencing. However, PMF-based protein identification relies on a single protein because mixed proteins can significantly complicate PMF analysis and may yield incorrect results. Therefore, for PMF identification with more than 2-3 protein samples, additional MS/MS-based protein identification is required to enhance the specificity of protein identification.
Peptide Mass Fingerprinting (PMF) is a proteomics technique that identifies proteins by analyzing the mass of peptide fragments produced from enzymatic digestion. Here are the detailed steps involved:
Protein Digestion: The target protein is digested using specific proteases, such as trypsin, which cleaves proteins at the carboxyl side of lysine and arginine residues. This controlled digestion produces a mixture of peptides with predictable masses. Other proteases like chymotrypsin or Glu-C can also be used to generate complementary peptide maps.
Mass Spectrometry Analysis: The peptide mixture is ionized using techniques such as Matrix-Assisted Laser Desorption/Ionization (MALDI) or Electrospray Ionization (ESI). These ionization methods convert peptides into gas-phase ions that are analyzed in a mass spectrometer, which measures the mass-to-charge (m/z) ratio of each peptide.
Peptide Mass Determination: The mass spectrometer generates a mass spectrum, a plot of the intensity of ions detected against their m/z ratios. Each peak in the spectrum corresponds to a peptide with a specific mass. High-resolution mass spectrometers, like time-of-flight (TOF) or Orbitrap, provide accurate mass measurements crucial for peptide identification.
Database Search: The experimentally obtained peptide masses are matched against theoretical masses calculated from protein sequence databases such as SwissProt or NCBInr. Software tools like Mascot, SEQUEST, and ProteinProspector compare the mass fingerprints to identify the protein by finding the best match between experimental and theoretical peptide masses.
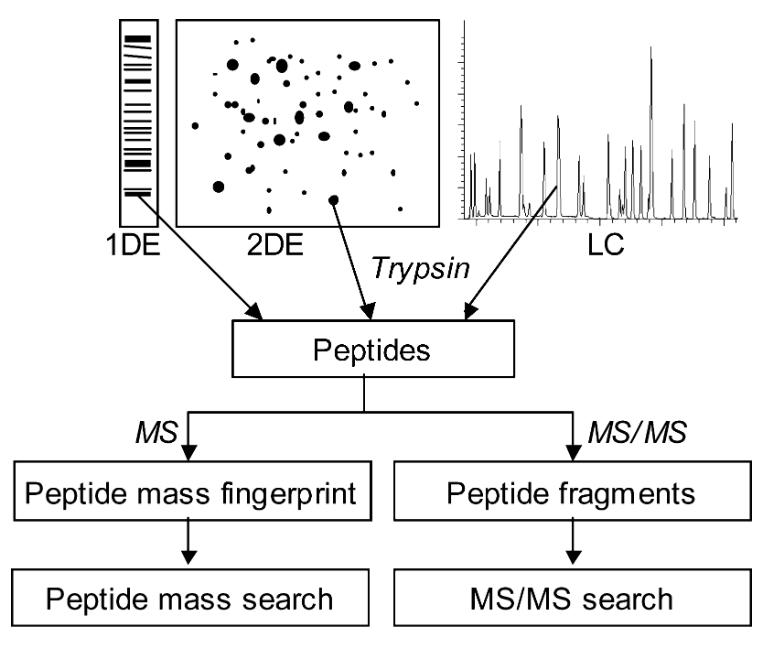 Common procedures to identify proteins by mass spectrometry. ( Thiede B, Höhenwarter W, Krah A, et al., 2005)
Common procedures to identify proteins by mass spectrometry. ( Thiede B, Höhenwarter W, Krah A, et al., 2005)
Peptide Mass Fingerprinting (PMF) works by first extracting and purifying proteins from biological samples. These proteins are then enzymatically digested, usually with trypsin, which cleaves the proteins at specific amino acid residues to produce a mixture of peptides. The peptide mixture is then ionized using techniques like Matrix-Assisted Laser Desorption/Ionization (MALDI) or Electrospray Ionization (ESI), which convert the peptides into gas-phase ions. These ions are analyzed by a mass spectrometer, which measures their mass-to-charge (m/z) ratios and generates a mass spectrum—a plot of ion intensity versus m/z ratio. Each peak in this spectrum corresponds to a peptide with a specific mass.
The mass spectrum produced forms a unique peptide mass fingerprint for the protein. This fingerprint is then compared to theoretical peptide masses derived from protein sequence databases using specialized software tools. The software matches the experimental peptide masses to the theoretical ones, identifying the protein based on the best fit. This process allows for high sensitivity and specificity in protein identification and can be applied to various samples, including cell lysates and tissue extracts, making PMF a powerful tool in proteomics for identifying proteins, studying post-translational modifications, and discovering disease biomarkers.
Peptide Mass Fingerprinting has numerous applications in various fields of biological research:
Protein Identification: PMF is a primary method for identifying unknown proteins in complex mixtures. For example, researchers can use PMF to identify proteins extracted from a gel band after electrophoresis, facilitating studies on protein expression and function.
Proteome Analysis: PMF is instrumental in large-scale proteomics projects aimed at cataloging all proteins expressed in a cell, tissue, or organism. By analyzing protein digests from different samples, PMF helps in mapping proteomes and understanding global protein expression patterns.
Post-Translational Modifications (PTMs): PMF can detect PTMs by identifying peptide masses that differ from the theoretical mass due to modifications. For example, a phosphorylated peptide will show a mass increase of 79.966 Da, allowing researchers to locate phosphorylation sites and study their regulatory roles in cellular processes.
Disease Biomarker Discovery: PMF is used to compare protein expression profiles between healthy and diseased tissues. Differences in peptide masses can indicate the presence of disease-specific proteins, aiding in the discovery of biomarkers for conditions like cancer, cardiovascular diseases, and neurodegenerative disorders.
Protein-Protein Interactions: PMF helps in identifying the components of protein complexes. By isolating protein complexes and analyzing their digests, researchers can determine the interacting partners and study the molecular mechanisms underlying cellular processes.
Peptide Mass Fingerprinting has several distinctive characteristics that contribute to its effectiveness in proteomics research:
High Sensitivity and Specificity: PMF can detect proteins present in low abundance due to the sensitivity of modern mass spectrometers. Specificity is achieved by the unique pattern of peptide masses generated for each protein, ensuring accurate identification even in complex mixtures.
Rapid and High-Throughput: PMF allows for the simultaneous analysis of multiple proteins, making it suitable for high-throughput studies. Automated sample preparation, digestion, and mass spectrometry analysis streamline the workflow, enabling the analysis of hundreds of samples in a relatively short time.
Minimal Sample Requirement: PMF requires only small amounts of protein samples, which is advantageous when working with limited or precious biological materials. This is particularly useful in clinical research, where sample availability can be a limiting factor.
Versatility: PMF is applicable to various sample types, including cell lysates, tissue extracts, and purified proteins. It can be combined with different proteases and mass spectrometry techniques, providing flexibility in experimental design and broad applicability across different research areas.
Quantitative Analysis: PMF can provide quantitative data on protein expression by comparing the intensities of peptide peaks in mass spectra. Techniques like isotope-coded affinity tags (ICAT) and tandem mass tags (TMT) enhance the quantitative capabilities of PMF, allowing for comparative analysis of protein expression under different conditions.
Complementary to Other Techniques: PMF complements other proteomic techniques such as tandem mass spectrometry (MS/MS), which provides sequence information for peptides, and liquid chromatography-mass spectrometry (LC-MS), which separates complex peptide mixtures before mass analysis. These combined approaches enhance protein identification and characterization.
Creative Peptides accepts samples of various types of dyed protein tapes, including Camas Bright Blue, Silver, Fluorescent, Zn/Cu2+, and others. We have no minimum requirements for protein content in the sample. The analytical method used by us is the most sensitive and reliable method for identifying protein tapes isolated from gels (including two-dimensional gels) in the world.
This service can be used to identify protein mixtures within 300 proteins. For example, protein complexes prepared by immunoprecipitation (antibodies should be cross-linked on a solid support), or protein complexes prepared by the Pull-down method. Compared to the identification of a single protein tape, this method can identify up to 300 proteins at a lower cost.
We can identify thousands of different proteins from a mixture sample and provide customers with complete protein data, to help customers obtain the complete protein data, especially low protein concentration, but very important.
Peptide Mass Fingerprinting (PMF) is an analytical technique used to identify proteins by comparing the mass of peptide fragments generated from protein digestion. By matching the mass spectra of peptides to theoretical values from protein databases, it allows for precise protein identification.
PMF involves enzymatic digestion of proteins into peptides, followed by mass spectrometry to measure their mass-to-charge ratios. The experimental peptide masses are then compared with theoretical values in protein databases to identify the protein. This technique is sensitive, accurate, and widely used in proteomics.
PMF is widely used for protein identification, proteome analysis, detecting post-translational modifications (PTMs), discovering disease biomarkers, and studying protein-protein interactions. It helps in mapping complex proteomes, understanding cellular processes, and identifying biomarkers for diseases like cancer and neurodegenerative disorders.
PMF offers high sensitivity, allowing for the detection of low-abundance proteins. It is also fast and can handle high-throughput analysis, making it suitable for large-scale studies. Additionally, it requires minimal sample amounts and can be applied to various sample types, including cell lysates, tissue extracts, and purified proteins.
Creative Peptides provides comprehensive PMF services, including protein identification from gel samples, protein mixtures, and complex protein data from low-concentration samples. Their advanced equipment ensures accurate and reliable results, making them a trusted partner for proteomic research.
Yes, PMF is effective in detecting PTMs like phosphorylation and glycosylation by identifying discrepancies between the theoretical and experimental peptide masses. This helps in studying the functional roles of modifications in proteins and their involvement in cellular processes.
PMF is a complementary technique to other methods like MS/MS and LC-MS. While MS/MS provides detailed sequence information, PMF focuses on identifying proteins based on peptide mass profiles. The combination of these methods enhances overall proteomic analysis and provides more comprehensive protein characterization.

References