Creative Peptides provides high-quality peptide array-based epitope mapping services to enhance your research. We harness the method's potential to offer a comprehensive view of the antibody response at an amino-acid level. Our services include peptide array design and synthesis, data processing and analysis, and expert consultation.
The peptide array technique is an innovative high-throughput method that enables the synthesis of thousands of peptides on a solid surface in a parallel manner. These peptides then serve as substrates for antibodies, revealing their antigenic determinants or epitopes. The exhaustive and precise nature of peptide arrays makes it a superior tool in epitope mapping over traditional methods as it allows for the simultaneous analysis of whole protein sequences.
The principle of the synthesis of peptide arrays and their use for antibody binding analysis is outlined in Fig. 1. A UV-light source combined with digital micromirrors is used to selectively activate small squares of the array, and amino acids with photo-labile protective groups are then added to the whole array. The amino acids will only be incorporated into the previously activated peptides, and through repeated cycles of activation and coupling, all unique peptides on the array can be synthesized in parallel. Because of the digital nature of the synthesis technology, array peptide sequences can be readily changed, enabling the synthesis of both custom and fixed-content array designs. After synthesis the arrays are incubated with the antibodies of interest, and this primary binding is detected with a fluorophore-conjugated secondary antibody. The arrays are scanned with a high-resolution microarray slide scanner, and the fluorescence intensities of the peptide features are evaluated. To cover the entire human proteome, based on the consensus coding sequences (CCDS) definition of the human proteome provided by NCBI, arrays were designed with 2.1 million overlapping 12-mer peptides with a six-amino-acid lateral shift.
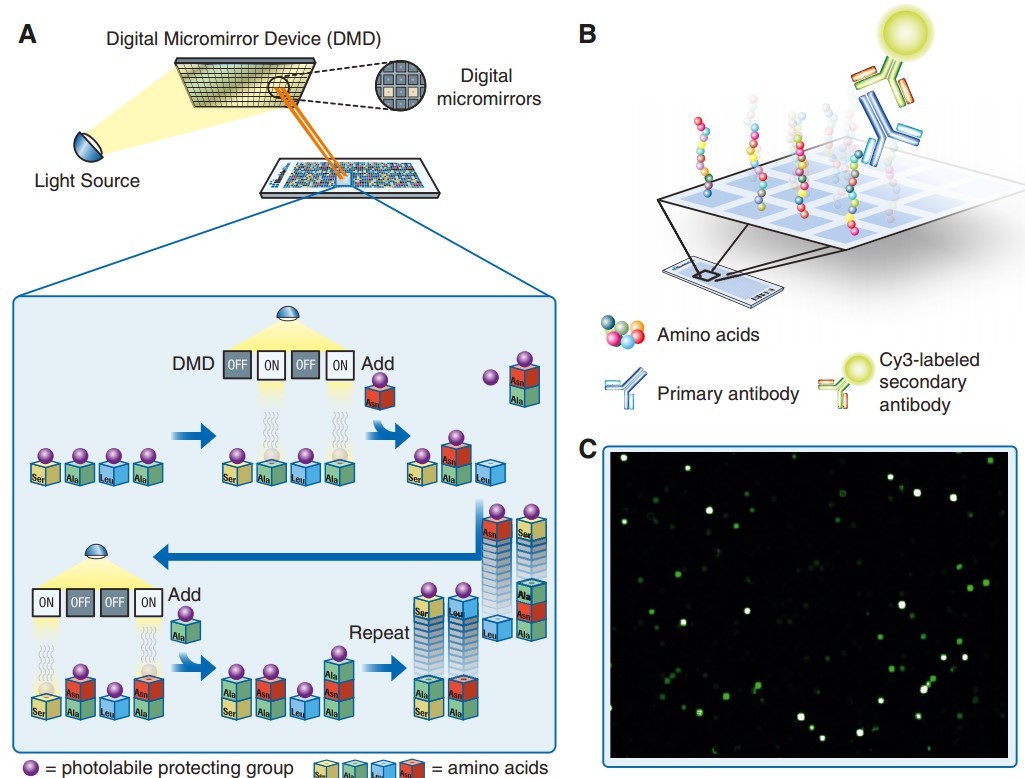 Fig. 1 The principle of in situ peptide array synthesis and subsequent antibody binding analysis. (Y Forsström, B., 2014)
Fig. 1 The principle of in situ peptide array synthesis and subsequent antibody binding analysis. (Y Forsström, B., 2014)
The technique used to define the antigenic determinants identified by individual antibodies is known as epitope mapping. This approach is vital because it elucidates the detailed interaction between an antigen and an antibody at an amino acid level. It allows for a comprehensive comprehension of the antibody-mediated immune response against specific diseases. A large number of methods have therefore been developed to determine the epitopes of antibodies, including mass spectrometry, solid phase libraries, and different display systems such as bacterial display and phage display. The most common method for epitope mapping involves the use of soluble and immobilized (tethered) peptide libraries, often in an array format, exemplified by the "Geysen Pepscan" method in which overlapping "tiled" peptides are synthesized and used for binding analysis. The tiled peptide approach can also be combined with alanine scans in which alanine substitutions are introduced into the synthetic peptides and the direct contribution of each amino acid can be investigated.
Peptide array-based epitope mapping involves the synthesis of overlapping peptides which cover the entire length of an antigen. This method allows for the high-throughput screening of multiple peptides and antibodies simultaneously, providing a comprehensive mapping of antigen-antibody interactions. It can also help in identifying immunodominant regions (areas of the antigen that elicit strong immune responses), which are crucial in the development of effective vaccines. Peptide array-based epitope mapping is a valuable tool in immunology research, aiding our understanding of the immune response, and facilitating the development of targeted medical therapies.
Epitope identification and epitope mapping can be performed by a biosynthetic peptide method. Compared to traditional phage library display, the biosynthetic peptide method can identify both the monoclonal antibody epitope and the polyclonal antibody epitope, and know all and fine linear B cell epitopes on the target protein. This provides conditions for revealing unknown epitopes of some pathogens, further preventing and controlling the occurrence of diseases. Epitope mapping can be used to detect the specificity of an immune response or the identification of different antibodies, as well as to determine the properties of a protein, such as the location of a protein, the source of a protein fragment, and the location of the protein in the cell.
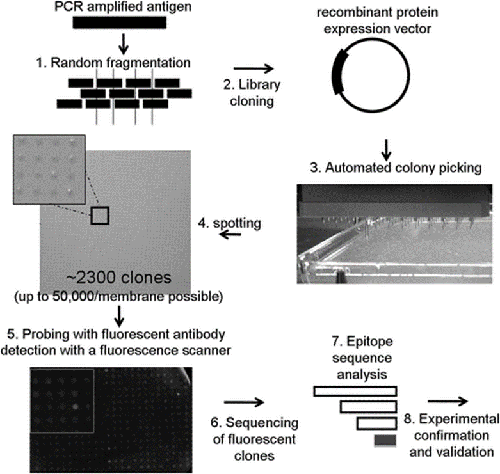 Fig. 2 Schematic of the epitope mapping procedure. (Maier, R. H., 2010)
Fig. 2 Schematic of the epitope mapping procedure. (Maier, R. H., 2010)
High-throughput: Peptide array-based epitope mapping allows the simultaneous screening of thousands of peptides. Thus, it can analyze a large amount of data in a relatively short period.
Cost-effective: The cost of peptide synthesis on a micro-scale is much lower compared to conventional peptide synthesis. Thus, this method is cost-effective for applications that require large scale screenings.
Sensitivity: This method can detect low affinity interactions, which might not be detected using other methods.
Versatility: It is capable of identifying both linear and conformational epitopes.
Speed: It is faster than other traditional methods of epitope mapping such as X-ray crystallography or NMR spectroscopy.
Easy to use: The procedure is relatively simple and does not require specialized skills or equipment.
Comprehensive: The entire protein sequence can be scanned for potential epitopes.
Quantifiable results: It provides direct, quantitative results. The intensity of signal in the array correlates with the strength of interaction between the protein and the peptide.
Low sample requirement: It requires less amount of sample compared to other methods.
Flexibility: Experiments can be easily designed to test specific hypotheses or examine particular regions of interest within the protein.
Using peptide array technology, researchers are able to perform high-throughput screening of target proteins and their effector binding sites, such as screening for intracellular kinase binding domains, and detecting changes in fingerprints of dysfunctional protein binding sites in the disease.
An epitope is a surface domain of an immunogen recognized by the corresponding antibody. Through multiple array technologies, all proteins of a virus or bacterium can be placed on a chip and reacted with sera from patients with different disease courses. The reaction site or binding domain is further screened to find specific epitopes for protecting antibodies. This epitope can be used for active or passive specific immune responses.
Researchers used focused-content peptide microarrays for epitope mapping of antibodies. The study focused on analyzing three polyclonal antibodies targeting recombinant fragments of HMGCR, HER2, and HYAL1 human proteins, which already had their three-dimensional structure determined. In this case, focused-content planar peptide arrays were designed with overlapping 12-mer peptides, covering the sequences of the protein fragments used for antibody generation, with a lateral shift of a single amino acid residue. The analysis was performed as technical replicates on two separate microarrays. The two technical replicates for the target HMGCR showed almost identical results with three distinct epitopes, two major epitopes and a minor epitope (green) close to the most N-terminal of the major epitopes. The three-dimensional structure of the native target showed that the three epitopes were parts of different structural elements, one α-helix, one β-pleated sheet, and one loop structure. The epitope mapping of the antibody toward the human epidermal growth factor receptor showed three distinct epitopes. The three-dimensional model of the native target showed that the epitopes were part of β-pleated sheets or loops. For the HYAL1 target, two major epitopes (green and cyan) were found using the microarray.
 Fig. 3 Epitope mapping of polyclonal antibodies toward three protein targets. (Y Forsström, B., 2014)
Fig. 3 Epitope mapping of polyclonal antibodies toward three protein targets. (Y Forsström, B., 2014)
Through the peptide array technology, hundreds of non-natural peptides are synthesized at one time to test the target protein, thereby promoting the discovery of the activity inhibitor or the competition conjugate. These inhibitors or competing conjugates can be used for subsequent development of candidate drugs for the treatment of disease. Drug development using the above-described polypeptide array technology is fast, flexible, and capable of achieving high throughput. processing.
Peptide array-based epitope mapping can aid in the detection of autoantibody signatures that are unique for specific diseases or cancers, providing a potential diagnostic tool.
It is used to identify allergen-specific IgE binding epitopes, which has implications for better understanding allergen-antibody interactions and development of novel diagnostics and therapeutics for allergies.
Creative Peptides provides comprehensive epitope mapping services. Simply send us your protein/antibody sample and we will perform an in-depth analysis of the sample and data results. We will use our excellent technology to provide Peptide Array-Based Epitope Mapping services to our global customers. Our epitope mapping integration services include:
Creative Peptides takes pride in being a trailblazer in offering cutting-edge peptide array epitope mapping services. Our technical expertise, supported by a highly-skilled research and development team, leverages these advanced technologies to develop pioneering therapeutic solutions.
Our company facilitates drug discovery by creating an extensive library of synthesized peptide sequences for array-based mapping, expediting the process of epitope identification. We validate the epitopes from the high-density arrays using in-house analytical tools for enhanced accuracy.
Moreover, we have cemented partnerships with research institutions globally to explore novel uses of peptide array-based epitope mapping in tackling challenging diseases. These collaborations help in optimizing our peptide synthesis processes, enhancing the quality, and broadening our array services for a diverse range of applications.
Peptide array-based epitope mapping service is an advanced technique used for determining the specific parts (epitopes) of an antigen that an antibody recognizes and binds to. This process involves using an array of peptide sequences to identify, analyze, and map the interactions between antibodies and antigens at a molecular level.
It works by synthesizing a large number of peptides onto a solid surface in a high-density array format. These peptides correspond to linear, continuous epitopes of the protein or antigen of interest. The array is then screened with the antibody in question and the bound antibodies are detected. The data obtained from this process is analyzed to identify which peptides are recognized by the antibody, thus indicating the location of the epitope on the protein of interest.
It has multiple applications in biomedical research, drug discovery, vaccine development, and diagnostics. It's particularly useful for understanding antibody-antigen interactions, identifying novel therapeutic targets, assessing immune responses, and developing and optimizing biologic therapeutics like vaccines and monoclonal antibodies.
The usual turnaround time varies depending on the complexity of the project, however, typically it takes about 2-4 weeks.
Peptide array-based epitope mapping service can be used to map epitopes from a wide range of antigens-from proteins and peptides, to complex biological structures like viruses and cells.
We ensure the accuracy of our services by using state-of-the-art lab equipment and validated protocols. Our experienced team of scientists also reviews the data to ensure accurate and reliable results.
To start a project, we require a clear understanding of the project parameters, including the protein sequence for the antigen of interest and a supply of the purified antibody. We may also require additional information depending on the specifics of the project.

References