The epitope substitution scan covers the stepwise substitution of all amino acid positions of the identified epitope with all 20 amino acids. In some cases, an epitope substitution scan can include selected unnatural amino acids, such as D-amino acids or citrulline. There are divided ito two types depending on the number of substituted amino acids used:
Epitope single substitution scanning encompasses a stepwise substitution of all amino acid positions of a given epitope or peptide with all 20 major amino acids and optionally other unnatural amino acids. The microarray we used included all amino acids, and finally, we can get a detailed amino acid map with conserved and variable amino acids.
In contrast to the stepwise exchange in epitope single-substitution scans, epitope double-substitution scanning enables high-throughput screening of epitope variants and mimotopes by exchanging two amino acid positions at a time. The epitope double substitution scan can output a detailed amino acid map. Therefore, epitope double-substitution scanning is a unique solution that can be used to develop and optimize peptide vaccines, peptide mimotopes or protein target binders.
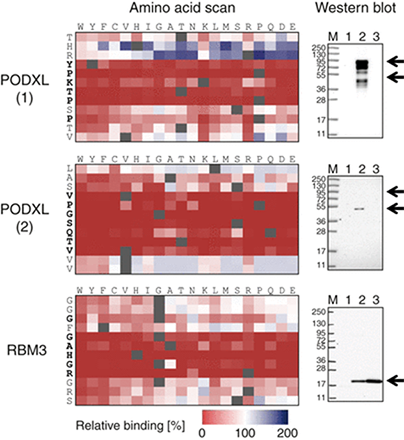 Fig.1 Full amino acid substitution scan and Western blot analysis of antibodies toward the human proteins PODXL and RBM3 (Forsstrm, et al. 2014)
Fig.1 Full amino acid substitution scan and Western blot analysis of antibodies toward the human proteins PODXL and RBM3 (Forsstrm, et al. 2014)
Samples that we can detect include: monoclonal or polyclonal antibodies, hybridoma supernatants, purified antibody components, serum, plasma, and the like.
Creative Peptides has developed a reliable epitope substitution scans in epitope mapping of peptide method. We can synthesize targeted peptide arrays according to customer requirements. We rigorously screen samples, perform precise analysis on results, and generate detailed reports. If there is a demand, please do not hesitate to contact us immediately, we will provide the best quality service.
An epitope substitution scan involves systematically replacing amino acids in a peptide sequence to analyze the impact on immune recognition and binding properties. This helps determine which amino acid positions are critical for immune activation and protein interaction.
In a single substitution scan, one amino acid is substituted at a time, allowing for precise mapping of each position's effect. A double substitution scan involves changing two amino acids at once, offering a faster way to identify key functional positions, particularly in high-throughput applications.
By identifying the essential amino acid residues within an epitope that interact with immune receptors, the substitution scan helps optimize peptides for vaccine development, ensuring stronger immune responses and more effective vaccines.
The method can be applied to a wide range of samples including monoclonal and polyclonal antibodies, hybridoma supernatants, purified antibody components, serum, and plasma, enabling comprehensive analysis of immune interactions.
By pinpointing immune-reactive regions within a protein, the scan allows modifications to reduce the immune system's recognition, thus minimizing adverse immune responses and improving the safety profile of protein-based drugs.
Creative Peptides offers highly reliable and customized epitope substitution scanning services, leveraging advanced technologies to ensure precise and timely results that contribute to the optimization of peptides and therapeutic development.
We synthesize custom peptide arrays based on specific requirements, then perform stepwise substitutions to evaluate each variant's immunoreactivity. The results are analyzed thoroughly, and we provide clients with detailed reports to guide further research or development efforts.

References